Professor, New Mexico Highlands University, NM
Website
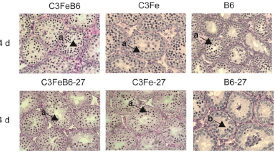
Mice carrying mutations that disrupt spermatogenesis provide valuable tools to identify and dissect pathways required for normal development. They provide clues to mechanisms of action when the process is disrupted leading to human infertility. Golgarepro27 is an ENU-induced allele of a golgin protein called GOLGA3 (i.e, Golgin-160) causing male-specific infertility. Mutant mice show defects during the first wave of spermatogenesis with most germ cells dying during mid-meiosis. GOLGA3 implicated in maintaining Golgi structure, protein targeting, and apoptosis signaling however its role in the testes is unknown. We want to identify GOLGA3-dependent pathways by looking at differences in transcriptome dynamics using RNASeq. A previous experiment identified 70 genes that were differentially expressed in mutant C57BL/6J-Golga3repro27 (B6-27) testes compared to C57BL/6J (B6) controls at 14 days of age. Pathway analysis identified genes primarily involved in spermatogenesis, meiosis and DNA repair. However, the germ cell populations were not equivalent as the B6-27 germ cells died earlier than what was expected. This RNASeq project examined mRNA gene expression in two B6-27 and two B6 control testes at 6 days of age, when the germ cell populations are the same. Students will use DNA subway and Discovery Environment tools to analyze raw data, explore gene structure/function relationships, and validate the data using real time PCR and western blotting. Identification of differentially expressed genes will provide clues to pathways that are altered in the absence of GOLGA3 and provide valuable clues its role in spermatogenesis and increase our understanding of genetic causes of human male infertility
NM-INBRE grants from NCRR (5p2RRP016480-12) and NIGMS (8P20GM103451) and a sequencing pilot award from the National Center for Genome Resources (NCGR) supported this research. Undergraduate student Devon Chisholm did the preliminary functional analysis for her senior research project (Chisholm et al., 2013). Students in my Cell and Molecular Biology course are conducting follow-up analysis using western blotting.
To assess the transcriptome dynamics involved in male infertility and identify potential genetic modifiers that exacerbate spermatogenic defects, RNASeq was used to compare wildtype and Golga3repro27 mutant mice in two well-established mouse strains: B6 (mt: C57BL/6J-Golga3repro27 (B627)) and C3Fe (mt: C3Fe.B6-Golga3repro27 (C327)). Testicular RNA (14 dpp) was isolated from 12 animals (3 for each genotype/strain), constructed into sequencing libraries with the IlluminaTruSeq protocol, and sequenced on the Illumina GAIIX platform, generating ~30,631,199 1x50nt reads/sample. Reads were aligned to the Mus musculus v10 (GRCm38) reference and uniquely aligned reads were binned and summated by gene to generate read count based gene expression estimates. Transcriptome wide analysis, including Principal Component Analysis and pairwise Euclidean distance calculations, clustered the B627 and B6 samples more tightly than the C327 and C3Fe samples. Differential gene expression analysis, as implemented by DESeq identified strain-specific gene expression signatures between mutant and wildtype in the B6 strain (91genes: 80 genes up regulated in B627) and C3Fe strain (127 genes: 125 genes down regulated in C327). Both genetic backgrounds identified 45 genes with differential expression between mutant and wildtype; intriguingly 44 of these genes (all but Golga3) showed divergent fold changes (Table 1). Between the wildtype and inbred strains, 1863 genes were differentially expressed. Preliminary functional analysis has identified genes involved in a number of pathways (Table 1). Quantitative PCR confirmed differential expression of two of these genes (Figure 2).
Table 1. Gene expression analysis
| Functional Annotation Clustering | Gene Symbol (Function or expression) | B627 > B6 (log change) | C327 > C3Fe (log change) |
|---|---|---|---|
| Golga3 (spermatogenesis, cargo trafficking, apoptosis) | -1.56 | -1.06 | |
| Defense Response Antimicrobial Proteins | C4bp(complement activation; stereocilia,Golgi complex, epithelia); Defb2 (extracellular component of epithelial cells); Defb6(dextracellular component, epididymis); Defb28 (extracellular component, testis-specific); Defb42 (active against gram-negative and positive bacteria, extracellular component); Wfdc15b (peptidase inhibitor, development of spermatocytes, spermatogenesis, extracellular domain,) | 2.36 – 3.51 | -2.73 – -4.12 |
| Cell adhesion | Cuzd1 (cell cycle, spermatogonia, spermatocytes and spermatids); Egfl6(cell cycle, epididymal growth factor); Cdh16(surface marker spermatogonial stem cells); Cldn3(tight junction of endothelial cells, Sertoli cells) | 1.09 – 2.51 | -1.13 – -2.36 |
| Cytoskeleton (intermediate filaments) | Krt5(spermatogonia); Krt8(epithelial integrity); Krt15(seminiferous tubules); Krt18(plasma membrane, spermatogenesis); Krt19 | 1.47 – 6.00 | -1.52 – -2.90 |
| DNA Transcription Factors Homeobox | Junb (germ cells, early spermatogonial development, chromatin remodeling); Pou3f3(HMG box domain binding, apoptosis, cell proliferation, nucleus); Ehf(early spermatogonial development, epithelial cells); Hoxa10(testis development); Hoxb8 (testis, spermatogenesis); Nr4a1(Sertoli cells) | 1.01 – 2.32 |
-1.18 – -2.65 |
| Peptidase and Protease Inhibitors | Serpina1f (spermatogenesis-associated); Spink8(spermatocytes, seminiferous tubules); Spink11 (apoptosis, sperm maturation); Wfdc15b(spermatocytes); Wfdc16(cystatin-related epididymis) | 1.74 – 2. 81 |
-2.91 – -4.70 |
The differential gene expression patterns observed in the existing dataset have raised a number of questions and warrant further research. Characterization of Golga3repro27 mutant mice on a mixed genetic background and on the C327 inbred strain showed significant germ cell loss occurring between 16 and 18 days of age, leading to the selection of 14 days of age for NGS analysis. However, subsequent histological analysis revealed that on the B6 inbred strain, germ cell loss occurred prior to 14 days and differences in the germ cell population could potentially explain the results. I am therefore proposing to extend gene expression analysis to a time point prior to the onset of meiosis on the more severe C57BL/6J inbred strain background. The new RNA-Seq dataset will allow the identification of gene pathways affected by the absence of GOLGA3. Follow-up analysis using quantitative PCR and protein analysis will allow us to explore our interest in genetic modifiers.
RNA-seq dataset to be provided at a later time