Assistant Professor of Biology, James Madison University
Website | Selected Works
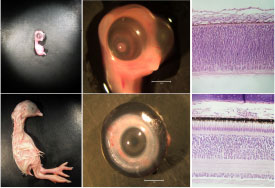
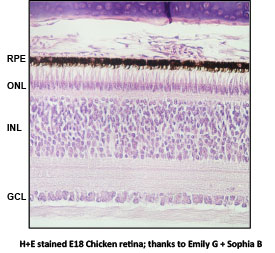
The chicken embryo (Gallus gallus) is a classic model system for studying developmental biology. My undergraduate research program in the Department of Biology at James Madison University uses the chicken embryo to study molecular mechanisms of retinal neurons during development. During chick development, embryonic day 8 (E8) retinas are packed with multipotent neuronal precursors on the cusp of differentiation into all retinal cell types. By E18 the retina is nearly fully mature with all major retinal cell types differentiated and expressing cell type-specific genes. Preliminary data from my lab demonstrates developmentally regulated fluctuations of the epigenetic modifier DNA methylation on several retina-specific genes. Methylation of retina-specific genes is inversely correlated with transcriptional activation during development. These observations have prompted me to use RNA-Seq to characterize global changes in gene expression between two critical developmental time points in the chicken retina.
Candidate genes identified in this dataset will be used to test our hypothesis that regulatory regions controlling retina-specific gene expression have an inverse correlation between levels of genomic DNA methylation and mRNA expression. Follow-up studies will investigate the role of DNA methylation and other epigenetic modifiers on developmentally regulated gene expression within retinal neurons. Due to the highly conserved nature of retinal programming among vertebrates, characterizing mechanisms controlling retinal development in chicken embryos will be valuable for gaining insight into conserved mechanisms of neuronal gene regulation as well as human retinal development and disease.
Whole retinas were harvested from embryonic day 8 (E8) and E18 developing chicken embryos. Total RNA was extracted using a Qiagen AllPrep Mini Kit. Duplicates were obtained from each time point. Illumina mRNA stranded TrueSeq RNA-Seq libraries were prepared from total RNA. 25-60 million paired-end reads were obtained per sample and mapped to the 2011 Galgal4 genome assembly. Differentially expressed genes between E8 and E18 retinas were calculated using the DNA Subway Green Line workflow and used for downstream analysis.
This RNA-Seq dataset was used to characterize global changes in gene expression between two critical developmental time points in the chicken retina.
E8 retina vs E18 retina Cuffdiff data (Download Excel)
As with most big data sets, the possibilities for follow-up studies are limited only by imagination. What can your students find with my data?
BIO480 Advanced Molecular Biology: An advanced biology lecture/lab course focusing on the history, principals & practices, and ethical considerations in the field of molecular biology. The course is taught every semester and serves 18 undergraduate students/section comprised of Biology, Biotechnology, and Chemistry majors. Lecture topics include a broad overview of DNA sequencing technologies and applications as well as student-led discussions based on primary literature and James Watson’s best selling book The Double Helix: A Personal Account of the Discovery of the Structure of DNA. The lab portion of the class consists of course-embedded research projects directly pertaining analysis and validation of RNA-seq data sets derived from retinal neurons. These activities involve hands on computer-based and wet bench molecular biology experiments.
BIO481 Genomics: An advanced biology lecture/lab course focusing on history, principals & practices, and ethical considerations pertaining to genomics and bioinformatics analysis. The course is taught once/year and serves 18 undergraduate students/section comprised of Biology, Biotechnology, and Chemistry majors. Lecture topics include in depth discussion of Sanger, Next-Gen, and 3rd Generation sequencing technologies and applications as well as student-led discussions based on primary literature and Matt Ridley’s best selling book Genome: The Autobiography of a Species In 23 Chapters. Lab activities consist of course-embedded research projects involving hands on computer-based and wet bench experiments analyzing and validating RNA-seq, ChIP-seq and other genomics data.