Professor, University of New Orleans, LA
Finite resource availability leads to trade-offs in several phenotypic traits that are key to fitness. The survival-fecundity trade-off is perhaps the best studied example and is caused by differential resource allocation toward one or the other, depending on a variety of factors. While life-history theory has been extremely useful in elucidating how resource allocation can result in adaptive responses to the environment, we know little about how traits critical to survival are subject to resource allocation trade-offs over an organism’s lifetime or how mechanisms that regulate these traits interact with other life-history traits.
In collaboration with Simon Lailvaux (University of New Orleans) and Jerry Husak (University of St. Thomas), using RNA-seq analysis, we aim to tease apart the proximate mechanisms underlying these life history trade-offs through transcriptomic analyses of key molecular pathways. Specifically, we hypothesize that dietary restrictions and exercise (sprint and endurance) training treatments will affect life history trade-offs through differential up- and down-regulation of specific molecular pathways. As a first step in understanding mechanisms underlying trade-offs between performance traits and other life-history traits, we have begun a pilot study to test the hypothesis that sprint training primarily alters the expression of IGF-1 in the liver whereas endurance training alters the HIF pathways in the kidneys of male green anoles Anolis lizards. In the Spring of 2015, we captured two A. carolinensis males from Washington park. One of these males was subject to sprint training by racing the animal 3 times per day on a 2.5 meter lizard racetrack with infrared beams spaced every 0.25 m to record speed, and the other for endurance training on a custom-built lizard treadmill set to 0.3 km/h once every other day. Training regimes were conducted for two weeks each prior to euthanasia and RNA isolation. Data from this study will allow us to compare transcriptome-wide differential gene expression among our two treatments in each of the pathways predicted below. We will also scan the transcriptome for evidence of differential gene expression in other candidate gene pathways.
At the completion of the experiment, animals were euthanized and the livers and kidneys were flash frozen and total RNA was extracted from both tissues using the RNAeasy minikit (Qiagen, CA). Quantity and quality of total RNA was assessed using the Experion microfluidic electrophoresis system (Biorad, CA). 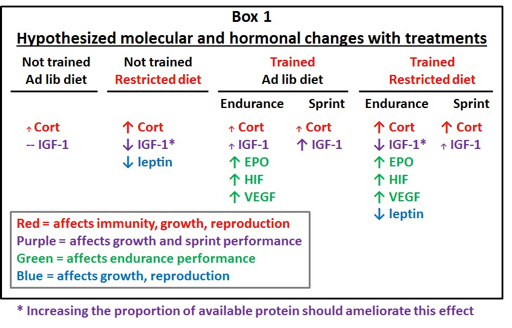
Materials are under development.
Materials are under development.
 Image courtesy of Jerry Husak
Image courtesy of Jerry Husak